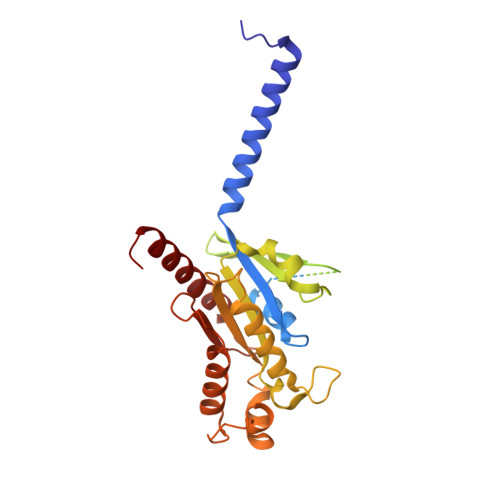
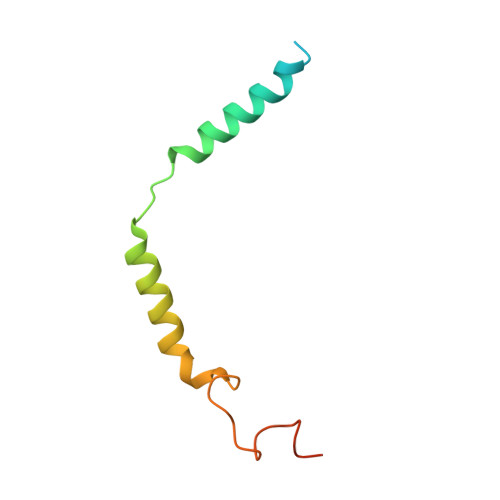
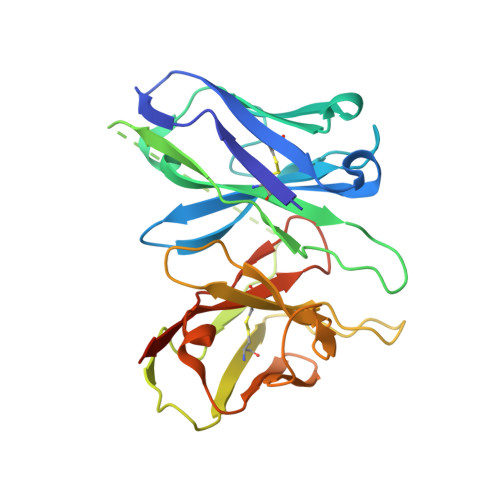
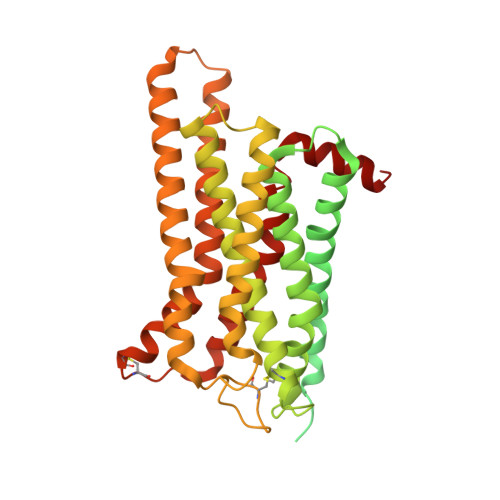
Structure of a D2 dopamine receptor-G-protein complex in a lipid membrane.
Yin, J., Chen, K.M., Clark, M.J., Hijazi, M., Kumari, P., Bai, X.C., Sunahara, R.K., Barth, P., Rosenbaum, D.M.(2020) Nature 584: 125-129
- PubMed: 32528175
- DOI: https://doi.org/10.1038/s41586-020-2379-5
- Primary Citation of Related Structures:
6VMS - PubMed Abstract:
The D2 dopamine receptor (DRD2) is a therapeutic target for Parkinson's disease 1 and antipsychotic drugs 2 . DRD2 is activated by the endogenous neurotransmitter dopamine and synthetic agonist drugs such as bromocriptine 3 , leading to stimulation of G i and inhibition of adenylyl cyclase. Here we used cryo-electron microscopy to elucidate the structure of an agonist-bound activated DRD2-G i complex reconstituted into a phospholipid membrane. The extracellular ligand-binding site of DRD2 is remodelled in response to agonist binding, with conformational changes in extracellular loop 2, transmembrane domain 5 (TM5), TM6 and TM7, propagating to opening of the intracellular G i -binding site. The DRD2-G i structure represents, to our knowledge, the first experimental model of a G-protein-coupled receptor-G-protein complex embedded in a phospholipid bilayer, which serves as a benchmark to validate the interactions seen in previous detergent-bound structures. The structure also reveals interactions that are unique to the membrane-embedded complex, including helix 8 burial in the inner leaflet, ordered lysine and arginine side chains in the membrane interfacial regions, and lipid anchoring of the G protein in the membrane. Our model of the activated DRD2 will help to inform the design of subtype-selective DRD2 ligands for multiple human central nervous system disorders.
Organizational Affiliation:
Department of Biophysics, The University of Texas Southwestern Medical Center, Dallas, TX, USA.